|
We have seen that protein aggregates complicate the lives of people who study protein folding in vitro
and who try to express human proteins in prokaryotes like E. Coli in vivo. Instead of viewing these aggregates as junk,
some now study them avidly. It turns out that these aggregates are not as non-specific as earlier believed. In addition, an
understanding of how and when they form will give us clues into the etiology and treatment of some of the most debilitating
and feared diseases. Much of this review is based on the following reference: Taubes, G. Misfolding the Way to
Disease, G. Science, 271, 1493-1495 (1996)
Clues Showing the Specificity of Aggregate Formation
- 1970's: It was shown that chymotrypsinogen could not be folded in vitro without
aggregates forming. An intermediate was presumed to have formed that if present in high concentration would aggregate irreversibly
instead of fold to the native state. Refolding of tryptophanase showed that it aggregated only with itself,
suggesting specificity.
- 1981: King, at MIT, found a single amino acid folding mutant in a viral protein. Both the normal and
mutant viral protein unfold at high temperature, but only the mutant would aggregate at high temperature, suggesting that
aggregation could be programmed into a gene
- mid 80's: The biotech industry, struggling to express growth hormone, found that a single
amino acid change in bovine growth hormone completely prevented aggregation without affecting correct folding.
Protein Aggregates and Disease
This knowledge of protein folding and aggregation was soon turn toward understanding several diseases
in which protein aggregates were observed which either initiated or were associated with disease. These protein aggregates
were termed "amyloid deposits" and seemed to be associated and perhaps causative of several neurodegenerative diseases. The
name amyloid was first used by a German pathologist, Rudolf Virchow, who in 1853 described waxy tissue deposits associated
with eosinophils (a type of immune cell). These deposits seemed to resemble starch (made of amylose and amylopecitn) so he
termed them amyloid. All known amyloid deposits are, however, composed of protein, not starch. It now appears that these disease
may be caused by improper protein folding and subsequent aggregation. Except in certain rare inherited diseases, the amyloid
deposits are composed of normal proteins (often called wild-type proteins) which seems to polymerize into fibrils. Sometimes,
in inherited conditions, or when mutations appear in a specific protein, the amyloid protein deposits consist of the mutant
protein. The proteins in these deposited fibers are composed predominantly of beta sheets which are perpendicular to the fiber
axis, while the unpolymerized protein usually has little beta sheet structure. Examples are given below:
- Familial amyloidotic polyneuropathy (FAP) - Affects 1/10,00 to 1/100,000 people. The protein
involved is called transthyrein, which normally exists in blood as a tetramer formed by association of 4
identical monomers. In mildly acid condition in vitro, the equilibrium between tetramer and monomer is shifted to monomer,
which can aggregate into fibrils. This aggregation could be promoted by possible transition to a molten globule (discussed
previously with lactalbumin) like state. This has secondary structure but loosely packed tertiary structure with more exposed
hydrophobes. If the concentration is high enough the molten globules aggregate. In people with the disease, mutations in the
protein destabilize the tetramer, pushing the equilibrium to the monomer, which presumably increases molten globule formation
and aggregation. Specifically, Val30Met and Leu55Pro mutations promote dissociation of the tetramer and formation of aggregates.
Conversely, Thr119Met inhibits tetramer dissociation. The aggregates deposit in heart, lungs, kidney, etc, leading to
death.
- Light Chain Amyloidosis; Light Chain Deposition Disease - The light chain protein is a
normal component of circulating antibody molecules. Mutants in the light chain cause a destabilization of the native state
to state similar to a molten globule, which then aggregates.
- Lysozyme amyloidosis - This protein, with extensive alpha-helix structure, is usually involved
in carbohydrate catabolism. Two mutants, Asp67His and another, Ile56Thr (normal amino acid/number in sequence/mutant amino
acid) destabliize the protein structure (as indicated by a decrease in the Tm of about 10 degrees C) to a molten-globule form,
which probably aggregates to fibrils characterized by extensive beta structure.
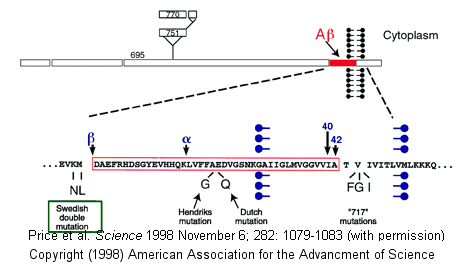
- Alzheimers-This disease involves a defect in a protein normally found in the membrane of
neurons. The protein, called beta-amyloid precursor protein (BAPP), is a transmembrane protein. A slightly truncated, soluble
form is also found secreted from cells and is found in extracellular fluid (such as cerebrospinal fluid and blood). The normal
function of these BAPP proteins is not yet clear. An endoprotease cleaves a small 40-42 amino acid fragment from this protein,
forming the amyloid beta (Ab) protein. It is this protein or a mutant
form of it which aggregates to form beta-sheet containing fibrils in Alzheimers disease. Several mutations in different proteins
have been linked to Alzheimers, but they all seem to increase production or deposition or both of the amyloid beta protein.
These deposited plaques are extracellular, and have been shown to cause neuronal damage. The are found in areas of the brain
required for memory and cognition. The BAPP gene is found on chromosome 21, the same chromosome which is present in an extra
copy (trisomy 21) in Downs Syndrome, whose symptoms include presenile dementia and amyloid plaques. Aggregate formation appears
to be driven by increase expression of BAPP and hence amyloid beta protein. In addition, some mutants may serve to destabilize
the amyloid beta protein, increasing its aggregation.
- Transmissible spongiform encephalopathies (TSEs) - Including
scrapie in sheep, bovine spongiform encephalopathy (mad cow disease), and in humans Creutzfeld-Jacob Disease (CJD), Fatal
Familial Insomnia (FFI),Gerstman-Straussler-Scheinker Syndrome, and Kuru (associated with cannibalism). In these fatal diseases,
the brain, on autopsy, resembles a sponge with holes. In contrast to the diseases above, these diseases can be transmitted from one animal to another, but typically not between species. (However, consider the controversy
with mad cow disease.) Also, the infectious agent can self-replicate in vivo. The logical conclusion is that a virus
(slow-acting) is the causative agent. However, the infectious agent survives radiation, heat, chemical agents, and enzymes
designed to kill viruses and their associated nucleic acids. Mathematical analyses suggested that the infectious agent in
such diseases could be nothing more than a protein. Stanley B. Prusiner in the 80's isolated just such a protein which he
named a prion, for proteinaceous infectious agent. Since then he and others have amassed
substantial evidence to support his contention. In October 1997 he was awarded the Nobel Prize in Medicine.
PRIONS
The normal cellular form of this protein, PrPc is highly conserved in mammals, and
is widely expressed in embryogenesis. Techniques exist to delete or make ineffectual genes in mice. When a mice knockout of
the PrPc (i.e. the gene for the protein was deleted in all cells) was made, the mice appeared normal. More recent
data suggests however, that these mice had altered circadian rhythms and sleep patterns, which suggest a possible link to
Fatal Familial Insomnia. The PrPc is a normal membrane protein in neurons. It is anchored to the membrane through
a glycosyl-phosphatidyl inositol link, with the protein chain on the outside of the neuronal plasma membrane. The PrPc
(without the PI link) is water soluble, protease sensitive, and consists of 42% alpha helix and 3% beta sheet.
 Chime Model: Prion Protein, Mad Cow Disease, and Mutations (made by Aaron Mohs and Amy Geissler Chime Model: Prion Protein, Mad Cow Disease, and Mutations (made by Aaron Mohs and Amy Geissler
 Chime Model: Prion Protein, Altegen, Inc Chime Model: Prion Protein, Altegen, Inc
The problem in the transmissible spongiform encephalopathies (TSE's) is that amyloid-like protein aggregates
form which appear to be neurotoxic. The protein found in the plaques (in cases other that those that are inherited) has the
same primary sequence as the PrPc but a different secondary and presumably tertiary structure. The protein found
in the plaques, called the PrPsc (the scrapie form of the the normal protein) is insoluble in aqueuous solution,
protease resistant, and has a high beta sheet content (43%) and lower alpha helix content (30%) than the normal version of
the protein PrPc.
Figure: Cartoon Models of PrPc and PrPsc
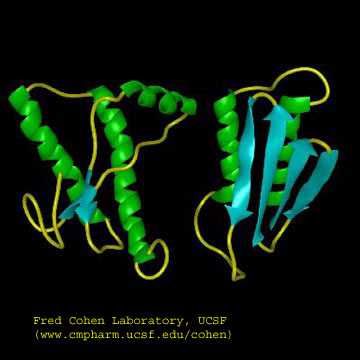
A genetic, inheritable form of disease also exists, in which a mutant form of the PrPc occurs,
whose normal structure is destabilized by the mutation. The aggregates caused by the mutant form of the disease are understandable
in light of the other diseases which we discussed above. The question is how does the normal PrPc form
PrPsc . Evidence shows that if radiolabeled PrP*c from scrapie free cells is added to unlabeled PrPsc
from scapie infected cells, the PrP*c is converted to PrP*sc! It appears that the PrPc
protein has two forms not that much different in energy, one composed of mostly alpha helix and the other of beta sheet. A
dimer of PrPc.PrPsc might form, which destabilizes the PrP*c causing a conformational
shift to the PrPsc form, which would then aggregate. Exposure to the PrPsc form would then catalyse
the conversion of normal PrPc to PrPsc . Hence, it would be transmissible by contact with just the PrPsc
form of the protein. Likewise species specificity could be explained if only dimers of PrPc.PrPsc
formed from proteins of the same species could occur. The inherited form of the disease would be explained since the mutant
form of the normal protein would more easily form the beta structure found in the aggregate.
It has recently been found that the very same mutation in PrPc, Asp178Asn
can cause two different diseases - CJD and FFI. Which disease you get depends on if you have 1 of two naturally occurring,
nonharmful variants at amino acid 129 of the normal PrPc gene. If you have a Met
at that position, and acquire the Asp178Asn mutation, you get CJD. If, on the other hand,
you have a Val at amino acid 129 and acquire the Asp178Asn
mutation, you get FFI. This disease was first observed in 1986
and has been reported in five families in the world. It occurs in the late 50's, equally in men and women. It is characterized
by a progressive loss of the ability to sleep and disrupted circadian rhythms. The brain shows neuronal loss. It is known
that amino acids 129 and 178 occur at the start of alpha helices, as predicted from propensity
calculations. Chronic exposure to micromolar levels of synthetic fragment 106-126 of PrPc kills hippocampal neurons.
This peptide also has the greatest tendency to aggregate synthetic PrPc peptides.
Kuru killed many members of the Fore tribe in New Guinea until the cannibalistic practice of eating dead
relatives was stopped. Analysis of the genes for the prion protein in the Fore tribe and other ethnic groups in the
world show two version differing by just one amino acid in all people (remember that a single gene is represented in
both maternal and paternal chromosomes. That these two forms exist through the world suggest that they have been selected
for by evolution and confer some biological advantage. People who have just one form of the protein are more susceptible
to the development of prion diseases. Mead and Collinge have shown that about 75% of older Fore women (who had lived
through cannibalistic practices) had two different prion genes, compared to about 15% of women from other ethnic groups.
This high percentage suggests that these women were protected from the disease, leading through natural selection to a high
percentage of heterozygotes in this defined population. The general presence of two forms of the prion gene (which probably
offers protection from prion disease) suggests that cannibalism might have been widespread in our early ancestors.
There appears to be one main difference between the formation of amyloid fibers from prion proteins and
others such as mutant lysozymes. If you add mutant lysozyme to normal lysozyme, the amyloid fibers contain only the mutant
protein. However, if you incubate mutant prion proteins with normal prions, the normal proteins become pathological.
QED - Protein Aggregates are not just test tube artifacts, but rather matters of life and death.
Recent work has shown the proteins considered to be completely harmless can generate misfolded intermediates
that aggregate to produce pre-fibril structures that are toxic to cells. This process is usually prevented in the cell
by interaction of nascent forms of the proteins with chaperones, which sequester exposed hydrophobic patches and prevent aggregation.
(Obviously prion proteins and the others mentioned above are exceptions). Amyloid fibers (characterized by subunits
with an abnormal amount of beta-structure) can be made from many different types of proteins as noted above. Is this
property specific to just a handful of proteins, or is it more common than expected from the limited examples noted so far?
The new studies show that when a bacterial protein HypF is incubated at pH 5.5 in the presence of trifluoroethanol, aggregates
(but not fibrils) form with enhanced beta structure. These aggregates slowly form into fibrils characteristic of amyloid
protein fibers. The early aggregates (before fibril formation) proved cytotoxic. Similar results were seen with
dimers and trimers (prefibril states) of the amyloid-b peptide released from
cultured neurons.
A diverse group of proteins that share not significant secondary or tertiary structure can form amyloid-like protein aggregates.
Even though their monomer forms share little in common, the insoluble amyloid aggregates have a common structure in which
the monomer in the aggregates has significant b-structure whose strands run perpendicular to the aggregate axis. Since
it has recently been shown that almost any protein, under the "right" set of conditions can form such aggregates, the stabilizing
feature of protein aggregates must be potentially found in any protein. Evidence suggests that is the presence of a
polypeptide backbone, which can form stable interstrand H-bonds in beta secondary structure, and not the side-chains, that
is the source of the common amyloid structure. In contrast, native, nonamyloid forms of normal proteins must arise through
specific interactions of unique side chain sequence and structure, which out competes nonspecific interactions among backbone
atoms found in amyloid structures. Nonspecific aggregation becomes more prevalent when buried hydrophobic side chains
and buried main chain atoms become more solvent exposed. Such exposure occurs when native proteins form intermediate
molten globule states when subjected to altered solvent conditions or when destabilizing mutants of the wild-type protein
arise. Some mutations may alter the cooperativity of folding which would increases the fraction of nonnative proteins
states. Other mutations that decrease the charge on the protein or increasing their hydrophobicity. In addition,
chemical modifications to proteins (such as oxidation or deamination) might destablize the native state, leading to
the formation of the molten globule state. Once formed, this state may aggregate through sequestering exposed side chain
hydrophobes or through inter-main chain H bond formation. Aggregate formation appears to proceed through the initial
formation of soluble units (which may or not be more toxic to cells than the final aggegate). Aggregates are kinetically
stable species. Since amyloid aggregates are cytotoxic and almost any protein can form them, albeit with different propensities,
nature, through evolutionary selection, has presumably disfavored proteins with high tendencies to form such aggregates.
|

