Any molecule that displays any of the catalytic motifs seen in the earlier guides (general acid/base catalysis,
electrostatic catalysis, nucleophilic catalysis, intramolecular catalysis, transition state stabilization) can be a catalyst.
So far we have examined only protein catalysts. These can fold to form unique 3D structures which can have active
sites with appropriate functional groups or nonprotein "cofactors" (metal ions, vitamin derivatives) that participate in catalysis.
There is nothing special about the ability of proteins to do this. It is now known that RNA, which can form complicated
secondary and tertiary structures as seen in the 3D image of the ribozyme from Tetrahymena thermophila, can as well.
Figure: ribozyme from Tetrahymena thermophila,

RNA molecules that act as enzymes are called ribozymes. This property of some RNA's was discovered
by Sidney Altman and Thomas Czech, who were awarded the Nobel Prize in Chemistry in 1989. In contrast to protein enzymes which are true catalysts in that they are used over again, this is an example
of a single use ribozyme. Other ribozymes are true catalysts and can carry out RNA slicing by transesterification
(splicesome) and peptidyl transfer (in ribosomes). The mechanisms of catalysis of the hepatitis delta virus
ribozyme include general acid/base catalysis.
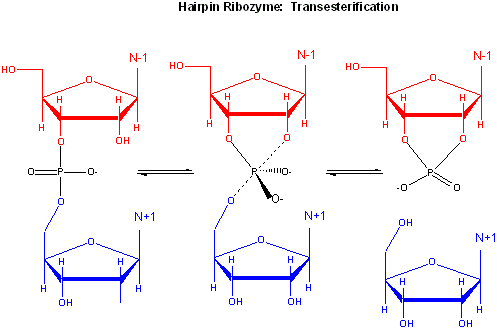
The hairpin ribozyme from satellite RNAs of plant viruses is 50 nucleotides long, and can cleave
itself internally, or , in a truncated form, can cleave other RNA strands in a transesterification reaction. The structure
consists of two domains, stem A required for binding (self or other RNA molecules) and stem B, required for catalysis. Self-cleavage in the hairpin ribozyme occurs in stem A between an A and G bases (which are splayed apart) when the 2' OH on the A attacks the phosphorous in the
phosphodiester bond connecting A and G to form an pentavalent intermediate.
Figure: Self-cleavage in the hairpin ribozyme
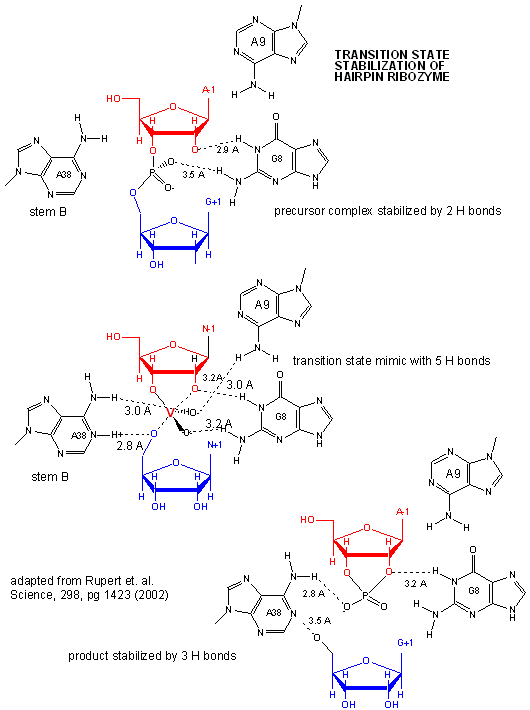
A recent study by Rupert et. al . (2002) shows that A38 in Stem B appears to be able to interact with
the products (the cleaved A now in the form of a cyclic phosphodiester with itself) and the departing G, and also with a transition
state pentavalent analog of the sessile A-G bond in which the phosphodiester linking A and G in the substrate is replaced
with a pentavalent vanadate bridge between A and G. However, A 38 does not appear to react with the sessile A -G groups
in the normal substrate, indicating that the main mechanism used by this ribozyme is transition state binding. Since
RNA molecules have fewer groups available for acid/base and electrostatic catalysis (compared to protein enzymes), ribozymes,
presumably the earliest type of biological catalyst, probably make more use of transition state binding as their predominant
mode of catalytic activity.
Recently, the crystal structure of a purple bacterium group I self-splicing intron (which catalyze the
removal of itself) interacting with both exons in a state prior to their ligation was determined (Adams, P. et al.).
The structure shows both exons in close proximity. Nucleophilic attack of the 3'OH of the 5' exon on a distorted phosphate
at the intron-3'-exon junction. Two metal ions reside on either side of the labile phosphodiester bond at
the intron-3'exon junction, and are held in place by 6 phosphates.
A novel use of ribozymes was recently reported by Winkler et. al. They discovered that the 3' end
of the mRNA of the gene glmS (from Gram-positive bacteria) which encodes an amidotransferase (catalyzing the formation
of glucosamine-6-phosphate from glutamine and fructose-6-phosphate) is a ribozyme. A glucosamine-6-phosphate binding
site in the ribozyme (3' end of the mRNA) binds this sugar, inducing autocleavage of the ribozyme. This inhibits, by
an uncertain mechanism, the formation of the amidotransferase from the remaining part of the mRNA. This mechanism of
regulation of gene expression through ribozyme activity might prove to be common.
Figure: Active Site of Hairpin Ribozyme: Transition State Binding
 Hairpin ribozyme with noncleavable substrate analog
Hairpin ribozyme with noncleavable substrate analog

